Poster: From single H&E to virtual immunohistochemical biomarker staining in the lung tumor microenvironment
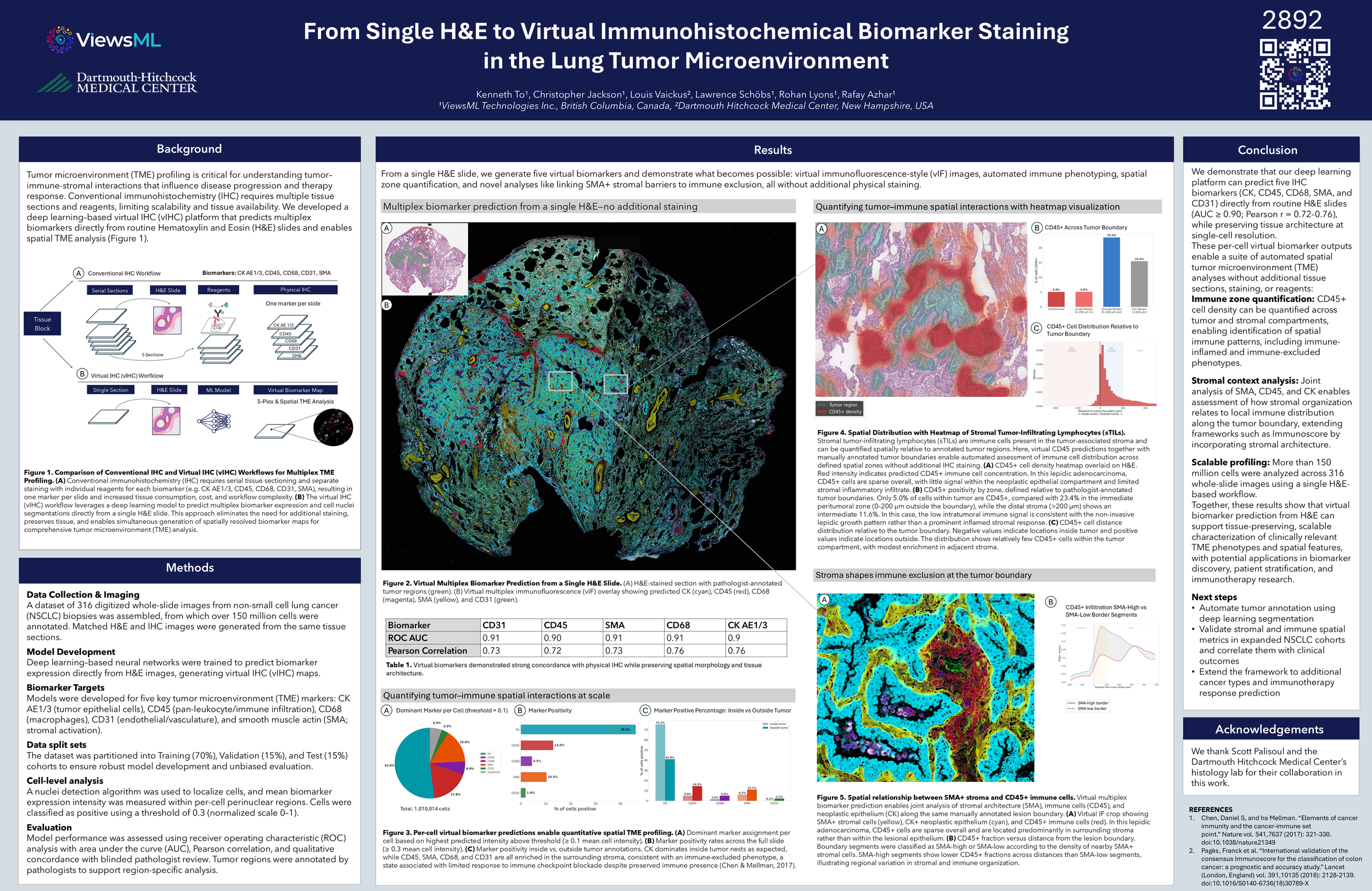
This poster describes a deep learning–based virtual immunohistochemistry (vIHC) platform that predicts multiplex biomarker expression directly from routine H&E slides to enable spatial tumor microenvironment (TME) profiling in non-small cell lung cancer (NSCLC). The approach replaces conventional multi-slide IHC workflows enabling high-fidelity TME characterization at single-cell resolution while conserving tissue.
Together, the data in this poster show:
Prediction of five key biomarkers (CK, CD45, CD68, CD31, SMA) from H&E slide
Quantitative spatial analysis of immune distribution, including tumor vs stromal zoning and immune-excluded phenotypes
Identification of stromal barriers (SMA+) associated with reduced immune infiltration at tumor boundaries
Scalable profiling across >150 million cells from 316 whole-slide images using a single H&E-based workflow